 
Deletions that target the open reading frame and result in frame-shifts, particularly early in the mRNA, are very likely to yield a non-functional protein sequence and to target the mRNA for nonsense-mediated decay due to premature stop codons in the new frame (Fig. A small, random deletion is often introduced at the repair site. In its simplest form, creation of a knockout line depends on the sgRNA-guided dsDNA cleavage by the Cas endonuclease, followed by non-homologous end joining (NHEJ) repair of the site by the cell. Thus, targeting of nearly any genomic sequence is possible with the introduction of an sgRNA and Cas9 into the cells of interest. Due to its particularly simple makeup, the type II CRISPR/Cas system of Streptococcus pyogenes has been adapted for genomic editing with great success: a single protein, Cas9, is required for crRNA binding and cleavage, and the RNA components have been engineered into a single guide RNA (sgRNA). Its transcription and processing yields small crRNAs that associate with and guide CRISPR-associated (cas) protein(s) to complementary DNA targets for endonucleolytic cleavage. Short (20–30 bp) sequence tags from the invaders are incorporated as spacers between direct repeats of a CRISPR locus. The CRISPR/Cas module is an endogenous adaptive immunity system commonly used by bacteria and archaea to counteract phage infection and introduction of plasmid DNA. Combinations of stable integration and modification with temporal control include the use of chemically inducible systems, such as tetracycline repression/activation, tamoxifen control of Cre-ER recombination, and more modular control of expression, localization and activity by small molecules. Often more experimentally desirable, stable genetic alteration can be achieved by integration of overexpression and shRNA constructs, or by genomic editing of the endogenous protein locus with transcription activator-like effector nucleases (TALENs), zinc-finger nucleases, and, more recently, by RNA-guided nucleases based on the clustered, regularly interspaced short palindromic repeats (CRISPR)/Cas9 system. Transient perturbations involve chemical inhibition (and activation) with small molecules, overexpression from non-integrating vectors, or knockdown by RNA interference. Manipulating protein levels and activities is a principal tool in understanding the functions and the relationships of these molecular components in cells. 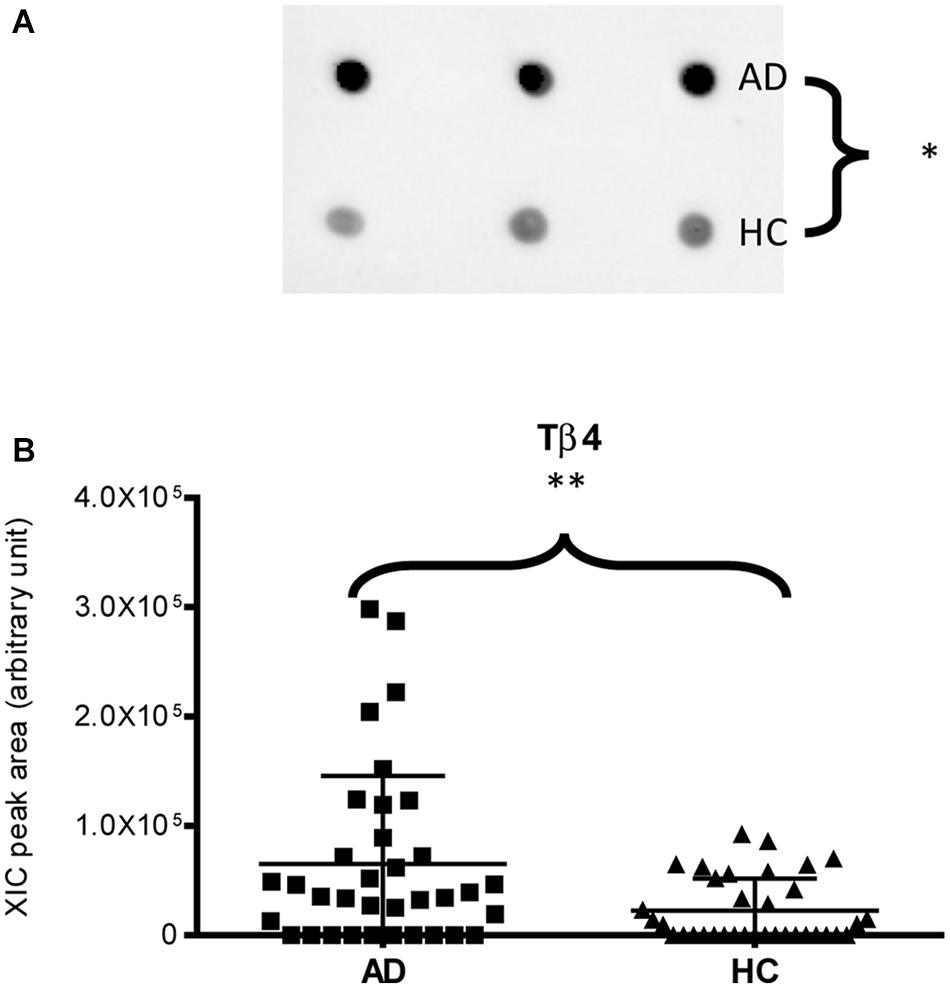
ConclusionsĬlonal screening for CRISPR/Cas9-mediated editing events using dot immunoblot is a straightforward and efficient approach that facilitates rapid generation of genomic mutants to study gene function. Genomic sequencing verified small deletions at the targeted locus. In total, 32 independent biallelic deletion lines out of 248 screened clones were isolated, and recovery of null mutants ranged from 6 to 36 % for the individual sgRNAs. Validation of knockout candidates by western blot indicated that the normalized protein abundances indicated by the dot blot serve as accurate predictors of deletion. To assess the technique, we probed clonal isolates of 293-TREx cells that were targeted with three separate sgRNAs against the HuR gene. We developed a procedure for rapid screening of clonal cell lines for the deletion of a protein of interest following CRISPR/Cas9 targeting in the absence of selective pressure based on dot immunoblots. However, streamlined approaches to establishing deletion and tagging mutants with minimal genomic perturbation are of interest in applying this methodology. Depending on the desired mutation, several experimental options exist in the isolation of clonal lines, such as selection with introduced markers, or screening by PCR amplification of genomic DNA. Targeted genomic editing using the CRISPR/Cas9 methodology has opened exciting new avenues in probing gene function in virtually any model system, including cultured mammalian cells. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
